Note
Click here to download the full example code
1D truncated unimodal example#
This example shows how to use lightkde.kde_1d with and without a limit and
how it compares to scipy.stats.gaussian_kde for a truncated unimodal distribution.
Import packages
import matplotlib.pyplot as plt
import numpy as np
from scipy.stats import gaussian_kde, norm
from lightkde import kde_1d
Generate synthetic data from a univariate normal distribution and truncate it:
np.random.seed(42)
sample = norm.rvs(size=2_000)
sample = sample[sample > 0]
Estimate kernel density using lightkde:
density_vec_without_x_min, x_vec_without_x_min = kde_1d(sample_vec=sample)
density_vec_with_x_min, x_vec_with_x_min = kde_1d(sample_vec=sample, x_min=0)
Estimate kernel density using scipy:
gkde = gaussian_kde(dataset=sample)
scipy_density_vec = gkde.evaluate(x_vec_without_x_min)
Plot the data against the kernel density estimates:
fig, ax = plt.subplots(figsize=(6, 4))
ax.plot(x_vec_with_x_min, density_vec_with_x_min, "--r", label="lightkde; with x_min=0")
ax.plot(
x_vec_without_x_min,
density_vec_without_x_min,
":r",
label="lightkde; without x_min",
)
ax.plot(
x_vec_without_x_min, scipy_density_vec, zorder=1, label="scipy.stats.gaussian_kde"
)
ax.hist(sample, bins=30, density=True, alpha=0.5, label="data")
ax.legend()
plt.show()
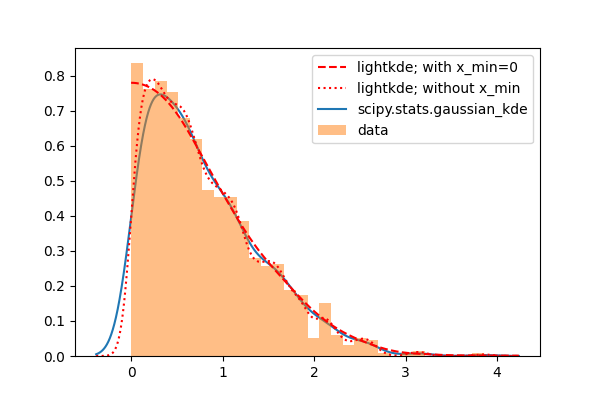
When x_min=0 is used, lightkde approximates well the histogram of the data.
Without this option it behaves similarly to the scipy method, both extend to a
region without data.
Total running time of the script: ( 0 minutes 1.908 seconds)